We often think the biggest challenge in neuroscience is generating better data. But after working across MRI, EEG, TMS, and biomarker studies, I’ve come to see something different:
The biggest challenge in translational research may not be scientific — but operational.
Neuroscience and clinical trials generate large multi-modal datasets. They span imaging, electrophysiology, blood biomarkers, cognitive testing, and clinical notes, across multiple visits. These types of studies are exciting for they can generate deep and novel insights about complex diseases, like Dementia, Depression, Motor Neuron Disease among others. But before insight, there is data.
A 1 hour neuroimaging (fMRI) visit may generate 10-50 gigabytes of raw data. This quickly becomes terabytes for the entire clinical trial. Yet the data and reports are commonly distributed across disconnected hardware and software systems that were never designed to talk to each other — each with their own formats and constraints.
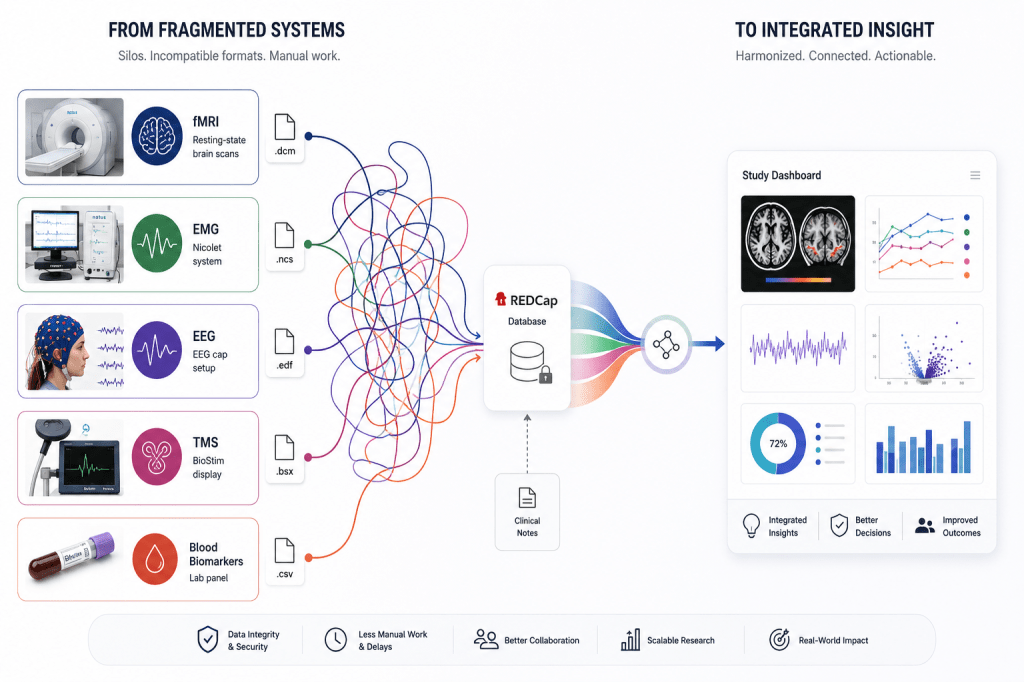
Better alignment across platforms could go a long way to creating meaningful insights faster.
Modern human-facing trials seek to advance personalized treatment regimes, understand complex clinical conditions and predict treatment-response.
The surprising part is not how much data exists. It is how fragmented the systems around it can be. Biological and clinical data are rarely captured in a unified system.
Some of the biggest bottlenecks in translational research may not be scientific — but operational.
To generate meaningful insights, these datasets must be brought together — often manually — across incompatible platforms and formats. Even when advanced computer programming tools exist, the final step of integration often depends on manual harmonisation.
What do I mean by manual? I mean the slow, old-fashioned process of clicking through clunky drop-down menus in grey software GUIs. Or meeting with team members to QC excel spreadsheets whos values were painstakingly copied by hand from faxed paper lab reports.
This is not a failure of any one system or team. In many cases, it reflects sound clinical governance. Let me explain.
Separating systems protects patient safety. It ensures compliance with tightly regulated ethical standards and allows the specialized teams to maintain data integrity by acting within their domain. The challenge emerges at the intersection between these systems — the spaces where information exists, but cannot easily flow. Hospital and research systems have strict firewalls, for good reason, which disable the sending of data over cloud or wired networks. It’s not uncommon to have to physically export imaging data onto a hospital-safe USB, then walk it down a long winding hallway that leads to your lab, and load it onto an analysis-enabled machine.
🧩 At times this can feel like trying to assemble a puzzle from a mix of LEGO, K’Nex, Meccano, jigsaw pieces, and custom-made Jenga blocks.
Valuable signals can become trapped behind incompatible formats, inconsistent metadata, and disconnected infrastructure.
These challenges are not abstract. They have real downstream consequences.
Poorly managed data integration can result in:
- slower identification of clinically relevant biomarkers
- reduced reproducibility across studies
- increased operational burden on research staff
- delays in translating findings into clinical insight
- difficulty scaling studies beyond small cohorts
Personnel, data infrastructure, cross-team coordination, clinical scheduling, governance requirements, and competing hospital priorities all shape what is possible. Not to mention the finite time and money. This is why I argue that research translation is not limited by a lack of scientific ambition. It is limited by the complexity of integrating real-world healthcare systems, people, and data.
And yet, this complexity may define the next frontier in neuroscience. 🧠 🤔
For years, the hurdle in human neuroscience was technology — how do we record signals from a brain that is floating, inaccessible inside the skull? Today, with non-invasive methods, like fMRI, EEG, and TMS, we can capture rich representations of brain activity in alive, awake, and even moving humans (pain free!). And even more may be possible with brain implants. The challenge will increasingly be how to connect the bitts.
Advances in open-source and AI programming tools have made integration more feasible.
Open-science frameworks such as BIDS and OpenNeuro, are helping standardise how neuroimaging data are structured and shared. Meanwhile, reproducible pipelines including fMRIPrep, MNE-Python, EEGLAB, FieldTrip and Clinica are enabling more scalable and transparent processing of imaging, electrophysiology and biomarker datasets. Put that together with AI-assisted coding, multi-level modelling packages shared on GitHub, global data systems like REDCap, and data visualisation tools such as Power BI, ggplot2 and Plotly, and the technology to process our big brain data is in our hands.
Applied to quality (non dubious) data with rigorous methods, these analytical tools can yield impactful discoveries with translational value.
I’m so excited to live at a time where it is possible to detect patterns in brain biomarkers at scales that even the Michael Shadlen‘s of the field might not have imagined a decade ago.
What’s clear to me now is that data + processing power is necessary, but not sufficient.
The future of translational neuroscience also depends on operational factors:
- shared standards (e.g., BIDS)
- interoperable systems that allow private sharing (e.g., EHRs and RedCap)
- thoughtful governance
- collaborative infrastructure
- and strong communication between teams
As a data scientist and biotech enthusiast, I am excited for the future of healthcare. It seems to me that we have the tools available to finally crack some of the biggest mysteries of the brain and body. There are already ventures operationalizing the power of integration, such as one called EveryCure.org, who are matching data on existing drugs with incurable diseases using AI tools. While I haven’t heard of similar projects for brain-based biomarkers (most of the current tools are in cell biology, genomics, and drug discovery, like AlphaFold, Evo, DeepBind, TOXRIC), I find this intersection highly promising.
As of now, I am convinced that some of the most meaningful breakthroughs in science will come not only from new discoveries — but from our ability to integrate data, operations and human-centered healthcare into the same conversation.
Real people are waiting for important and life saving treatments on the other side.
Curious how others working in translational research, health data, or clinical trials are seeing this problem.
